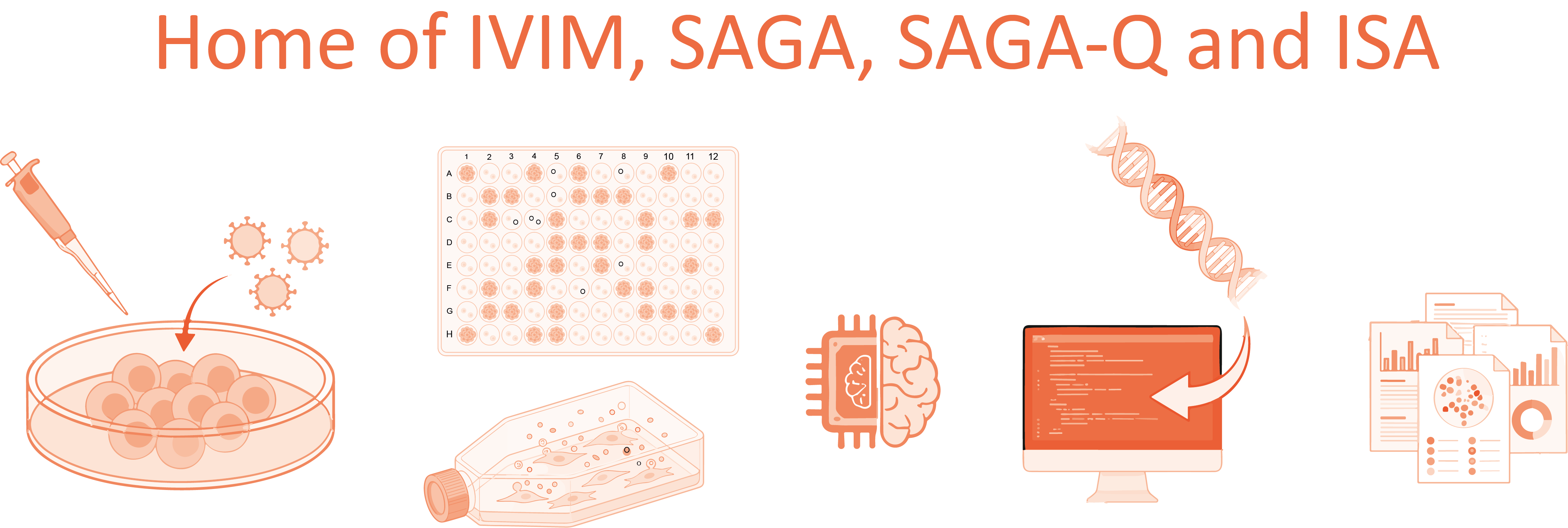
Check out the available assays:
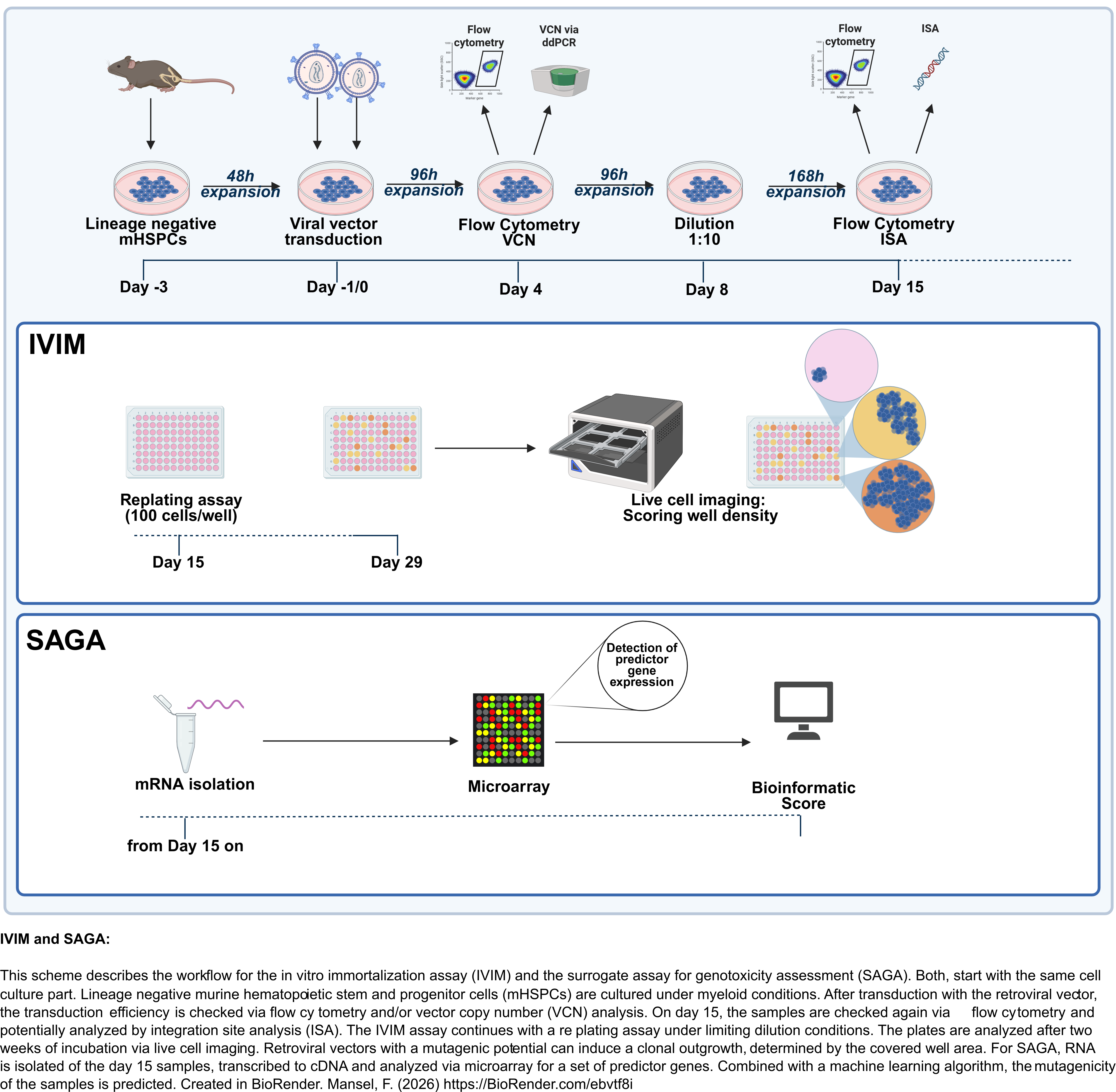
- IVIM (In Vitro Immortalization Assay): This is a biological "stress test" where mouse bone marrow cells are transduced with a viral vector and grown in culture. If a vector is genotoxic, it can cause "insertional mutagenesis"—mutating the DNA in a way that makes the cells immortal—leading to rapid, uncontrolled cell growth that signals a high risk of causing leukemia.
Read the initial article on the discovery of the IVIM here: Modlich, Blood 2006
Dive deeper into the assay with a paper on 18 years of experience with the assay: Bastone, Human Gene Therapy 2025
- SAGA (Surrogate Assay for Genotoxicity Assessment): SAGA is a sophisticated upgrade that uses a specific 11-gene expression signature to identify dangerous vectors. Instead of waiting weeks to see if cells grow uncontrollably (like in IVIM), SAGA uses machine learning to detect the molecular "warning signs" of cancer-causing changes almost immediately, offering much higher sensitivity and speed.
Read the full article describing the road to SAGA: Schwarzer, Molecular Therapy 2021
Here is the official press release from MHH describing the benefits of SAGA.
- Enhancing Vector Safety: Together, these tools allow researchers to screen out high-risk gene therapy vectors early in development. By identifying which vector designs are most likely to trigger harmful mutations, they ensure that only the safest configurations proceed to clinical trials, significantly reducing the risk of cancer for patients.
- Contact us for further information at genotoxicity@mh-hannover.de
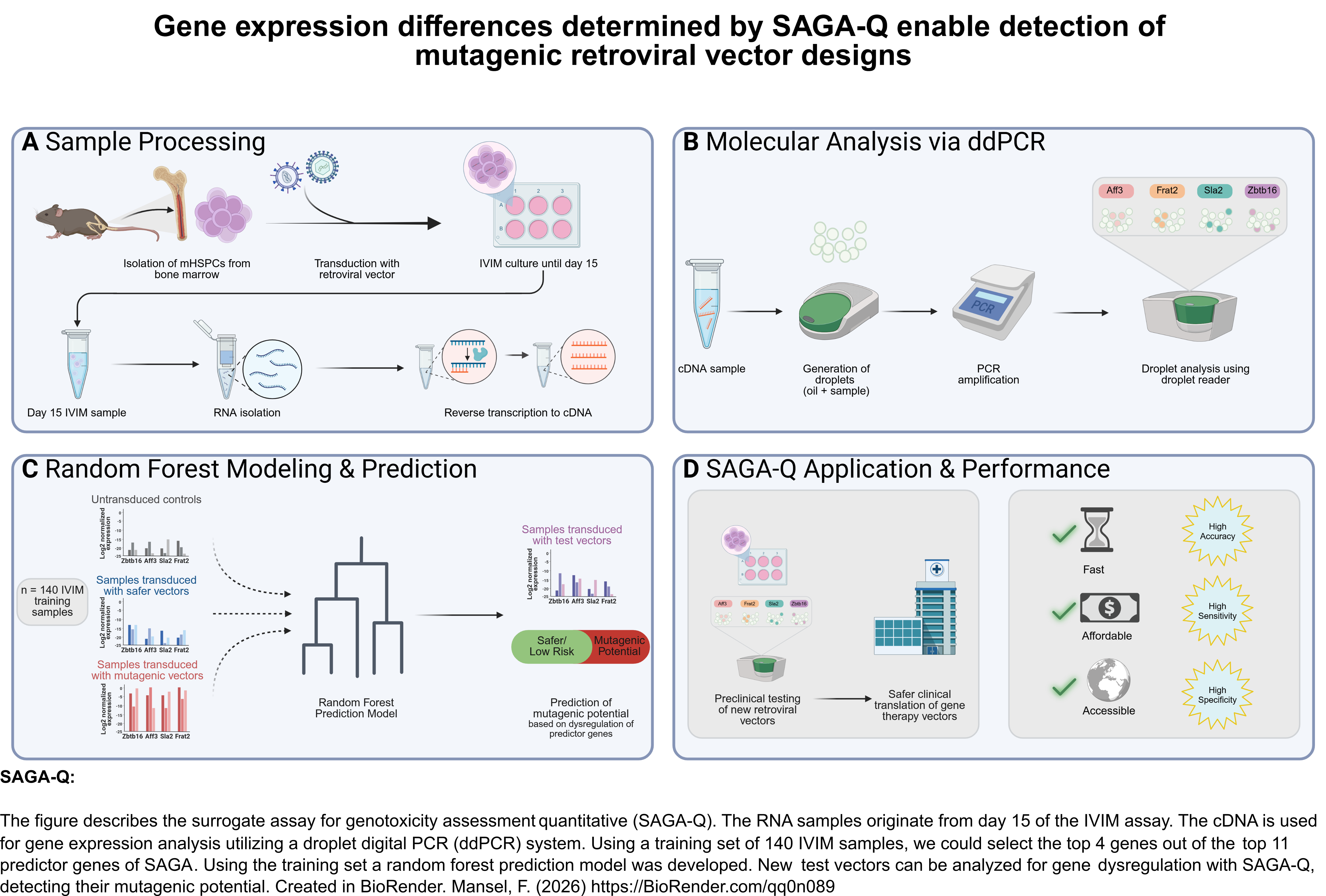
SAGA-Q improves upon the original SAGA method by employing a Digital Droplet PCR (ddPCR)-based readout, which streamlines genotoxicity assessment and lowers the barrier for clinical translation. By utilizing specific gene expression differences, SAGA-Q identifies mutagenic retroviral vectors more precisely and cost-effectively than the high-throughput sequencing or microarrays required by SAGA. This technological advancement enables faster, more accessible safety evaluation for gene therapy products prior to clinical trials. Read the full study here: SAGA-Q paper by Friederike Mansel.
Contact us at: genotoxicity@mh-hannover.de
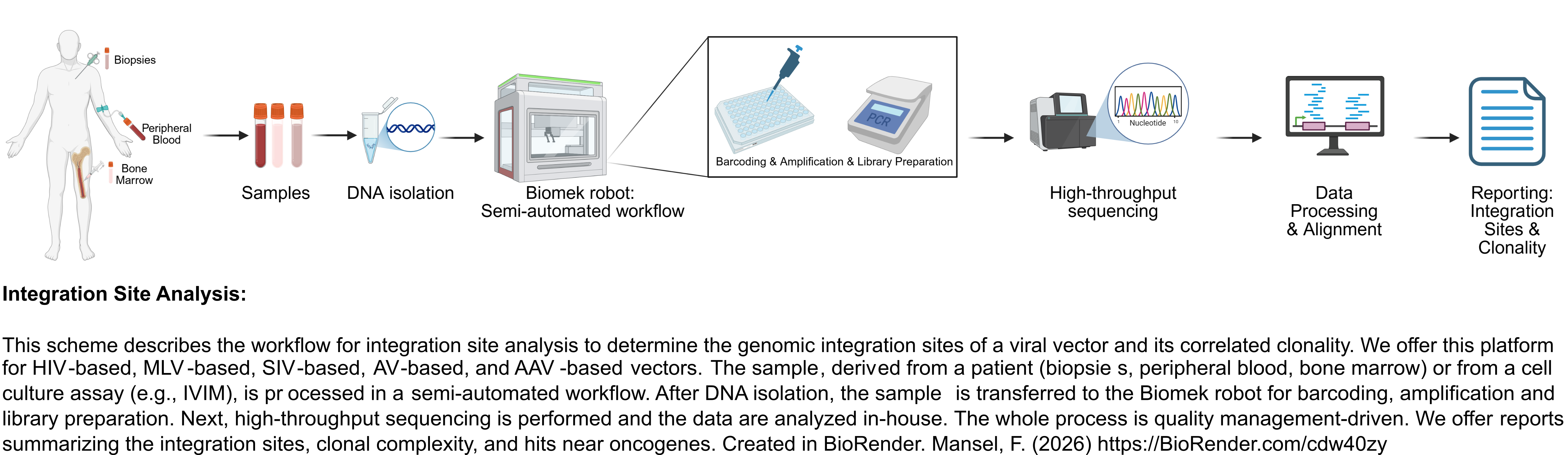
We offer a state-of-the-art integration site analysis platform to monitor patients in clinical trials. We also perform preclinical studies to determine vector safety and analyze causal relationships between proliferative neoplasms post gene therapy.
Check out high ranking publications we supported:
Lentiviral Gene Therapy for Severe Leukocyte Adhesion Deficiency Type 1 - The New England Journal of Medicine - This therapy, also known as KRESLADI, recently received FDA market approval.
Contact us for further information: genotoxicity@mh-hannover.de
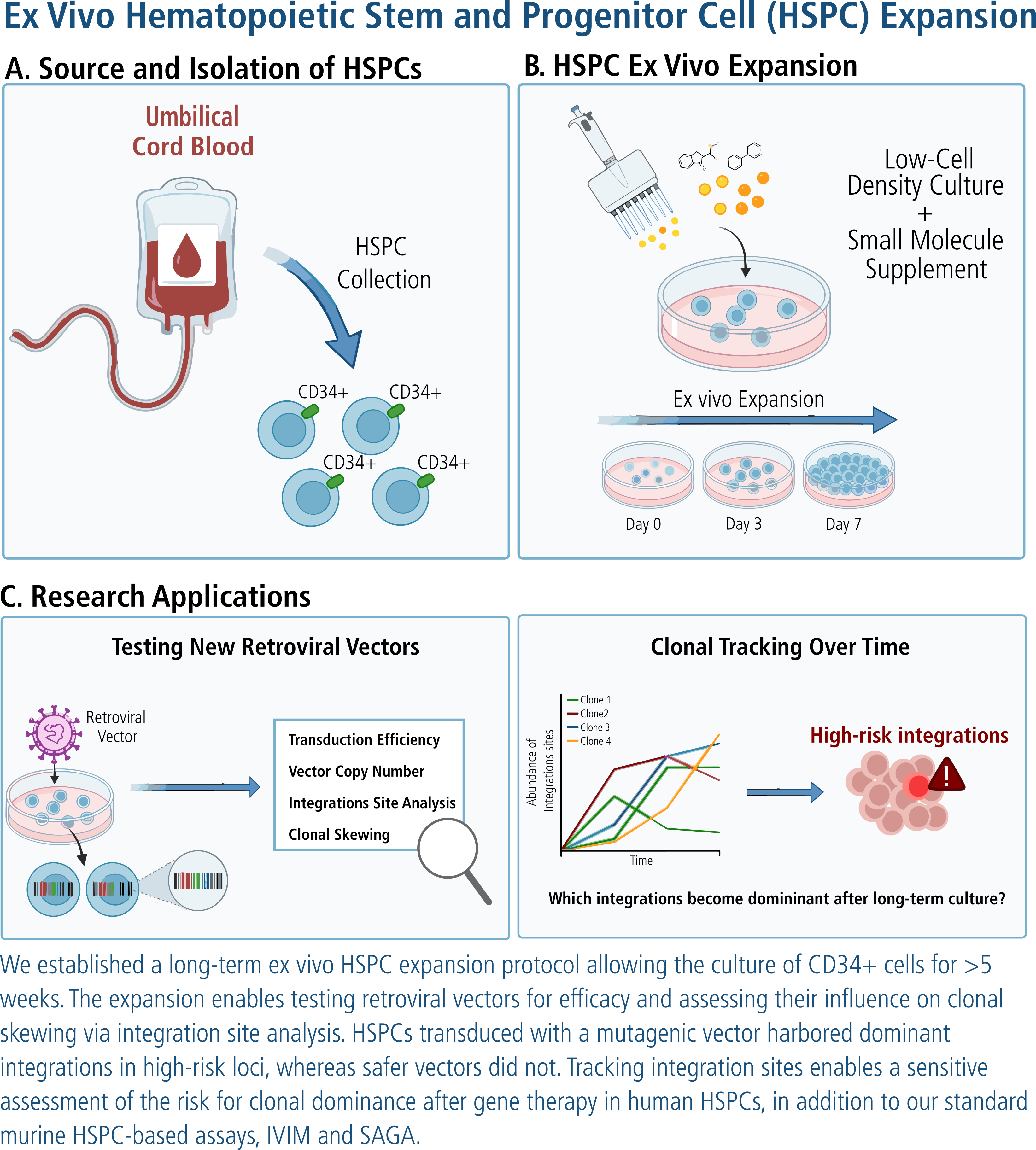
Precision Tracking of Clonal Evolution: See exactly how your vector behaves over time! By enabling the outgrowth of specific clonal integration sites, this assay allows you to monitor clonal dominance and Retroviral Integration Sites (RIS) with high resolution. It captures the molecular "red flags"—such as integrations in high-risk loci like MEIS1 or SUSD6—before they ever reach a patient.
Read the full study here: Clonal tracking paper by Jenni Fleischauer
This paper received the DG-GT Paper-of-the-Quarter Award Q1 2026.
Contact us for more information: genotoxicity@mh-hannover.de
For further information, contact us at: genotoxicity@mh-hannover.de
About us
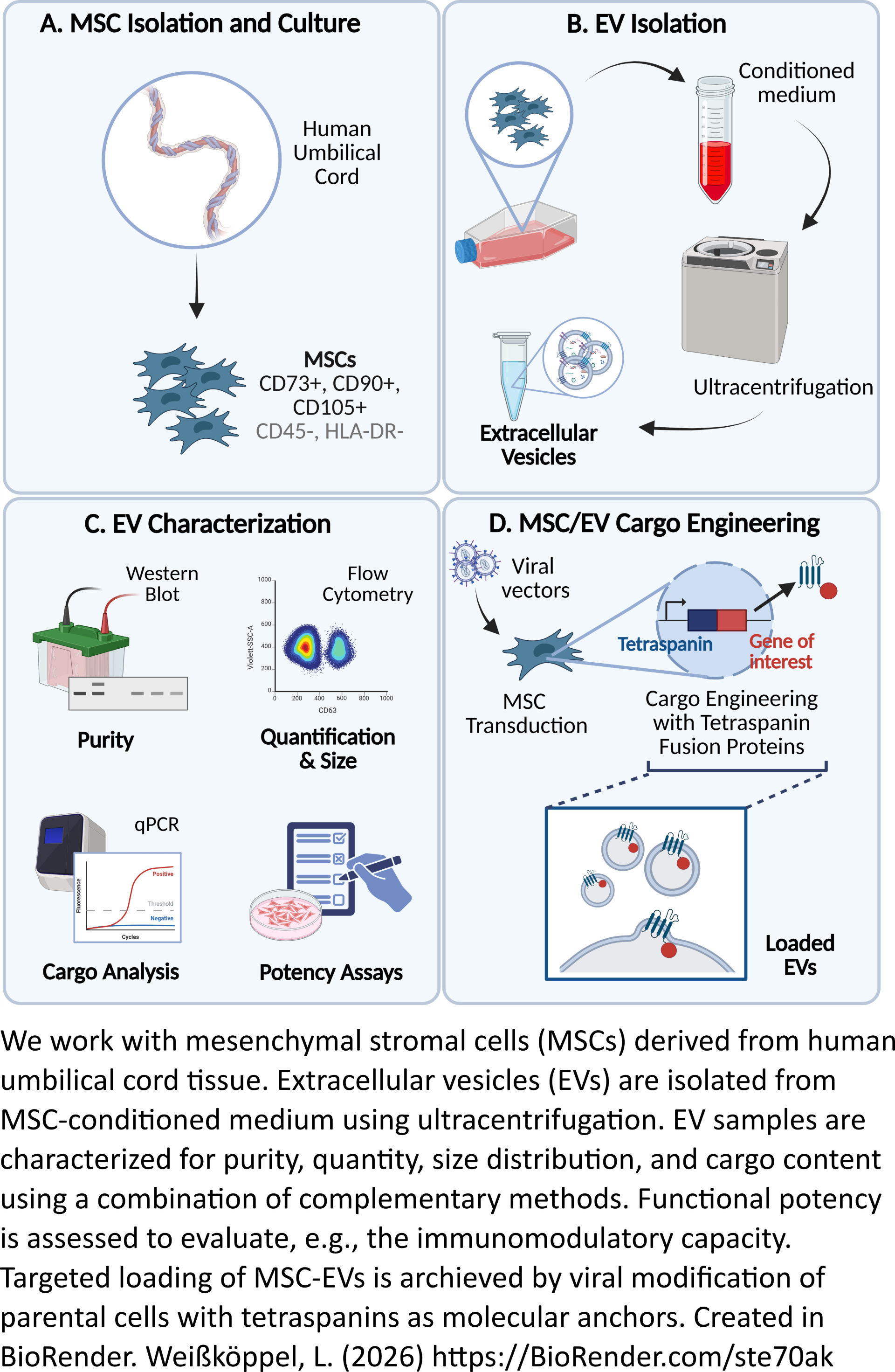
Our primary research goal is to establish innovative tools for preclinical risk assessment of viral vectors to minimize side effects like genotoxicity early in development. We deploy cutting-edge methods, including machine learning, to conduct safety and gene expression analyses of modified cells. A major part of our work is dedicated to the characterization and optimization of hematopoietic stem and progenitor cells and mesenchymal stromal cells to improve their therapeutic potential and post-transplantation survival rates. By linking basic biological research with translational approaches, we pave the way for clinically safe, next-generation cell and gene therapies.

From left to right: Philipp John-Neek, Friederike Mansel, Michael Rothe, Violetta Dziadek, Luisa Weißköppel, Jenni Fleischauer, Maike Hagedorn, Teng Cheong Ha. Missing on the picture but part of our team: Uta Limper.